Odisha State Board CHSE Odisha Class 12 Biology Important Questions Chapter 7 Molecular Basis of Inheritance Important Questions and Answers.
CHSE Odisha 12th Class Biology Important Questions Chapter 7 Molecular Basis of Inheritance
Molecular Basis of Inheritance Class 12 Important Questions CHSE Odisha
Very Short Answer Type Questions
Choose the correct option
Question 1.
In a DNA strand, the nucleotides are linked together by
(a) glycosidic bonds
(b) phosphodiester bonds
(c) peptide bonds
(d) hydrogen bonds
Answer:
(b) phosphodiester bonds
Question 2.
In DNA double helix, thymine is paired with …………… .
(a) guanine
(b) uracil
(c) cytosine
(d) adenine
Answer:
(d) adenine
Question 3.
Semiconservative mode of replication of DNA was proved by
(a) Hershey and Chase
(b) Griffith
(c) Watson and Crick
(d) Meselson and Stahl
Answer:
(d) Meselson and Stahl
Question 4.
Discontinuous synthesis of DNA occurs in one strand, because
(a) DNA molecule being synthesised is very long
(b) DNA dependent DNA polymerase catalyses polymerisation only in one direction (5′ → 3′)
(c) it is a more efficient process
(d) DNA ligase has to have a role
Answer:
(b) DNA dependent DNA polymerase catalyses polymerisation only in one direction (5′ → 3′)
![]()
Question 5.
The enzyme not associated with DNA replication is
(a) polymerase
(b) helicase
(c) topoisomerase
(d) transcriptase
Answer:
(d) transcriptase
Question 6.
Which is the enzyme used for joining the fragments of DNA?
(a) Ligase
(b) Polymerase
(c) Endonuclease
(d) Transferase
Answer:
(a) Ligase
Question 7.
To form a continuous DNA molecule, the enzyme ………… joins Okazaki fragments.
(a) primase
(b) polymerase
(c) helicase
(d) ligase
Answer:
(d) ligase
Question 8.
DNA ligase is commonly known as ……….. .
(a) molecular scissor
(b) molecular marker
(c) molecular probe
(d) milecular glue
Answer:
(d) molecular glue
Question 9.
In eukaryotic cells, the RNA transcribed from DNA is called ………….. .
(a) rRNA
(b) cistron
(c) cDNA
(d) heterogenous mRNA
Answer:
(d) heterogenous mRNA
Question 10.
At 5′ end of a polynucleotide chain
(a) H-bond is present
(b) -OH group is attached
(c) PO–4 group is attached
(d) pentose sugar is attached
Answer:
(c) PO–4 group is attached
Question 11.
In which one of the following, double-stranded RNA is present?
(a) Bacteria
(b) Chloroplast
(c) Mitochondria
(d) Reovirus
Answer:
(d) Reovirus
Question 12.
DNA replication is
(a) semiconservative, directional and continuous
(b) semiconservative, bidirectional
(c) semiconservative and semidiscontinuous
(d) only semiconservative
Answer:
(c) semiconservative and semidiscontinuous
Question 13.
A double-stranded RNA segment has 120 adenine and 120 cytosine bases. The total number of nucleotides present in the segment is
(a) 120
(b) 240
(c) 60
(d) 480
Answer:
(b) 240
Question 14.
Termination codon which stops, further addition of amino acids to the polypeptide chain is
(a) AAU
(b) GUG
(c) AUG
(d) UAG
Answer:
(d) UAG
Question 15.
Which one is not a non-sense codon?
(a) UAA
(b) UGA
(c) UCA
(d) UAG
Answer:
(c) UCA
Question 16.
A phenomenon where the third base of tRNA at its 5′ end can pair with a non-complementary base ofmRNAis called
(a) universality
(b) colinearity
(c) degenerency
(d) wobbling
Answer:
(d) wobbling
Question 17.
Translation is the synthesis of
(a) DNA from a mRNA template
(b) protein from a mRNA template
(c) RNA from a mRNA template
(d) RNA from a DNA template
Answer:
(b) protein from a OTRNA template
Question 18.
The peptide bonds are present between
(a) nucleic acids
(b) organic acids
(c) fatty acids
(d) amino acids
Answer:
(d) amino acids
Question 19.
Gene which is responsible for the synthesis of a polypeptide chain is called
(a) operator gene
(b) regulatory gene
(c) promoter gene
(d) structural gene
Answer:
(d) structural gene
Question 20.
Repressor protein is produced by
(a) regulator gene
(b) operator gene
(c) structural gene
(d) terminator gene
Answer:
(a) regulator gene
Question 21.
In split genes, the coding sequences are called
(a) cistrons
(b) operons
(c) exons
(d) introns
Answer:
(c) exons
Question 22.
The non-sense codons
(a) have no role in biological systems
(b) act as terminators during protein synthesis
(c) are of little value in transcription
(d) have a poor role in transcription
Answer:
(b) act as terminators during protein synthesis
Question 23.
If a cell is treated with a chemical that blocks nucleic acid synthesis, which of the following processes is the most likely one to be affected first?
(a) DNA replication
(b) tRNA synthesis
(c) mRNA synthesis
(d) Protein synthesis
Answer:
(a) DNA replication
Question 24.
Aminoacyl synthetase takes part in
(a) attachment of mRNA to 30S ribosome
(b) transfer of activated amino acids to tRNA
(c) activation of amino acid
(d) hydrolysis of ATP to AMP
Answer:
(c) activation of amino acid
Correct the statements, if required, by changing the underlined word(s)
Question 1.
Watson and Griffith proposed the double helical structure of DNA.
Answer:
Watson and Crick.
Question 2.
The helical turns are left-handed in Z-DNA.
Answer:
It is correct.
Question 3.
Okazaki fragments are formed on both leading and lagging strand.
Answer:
Okazaki fragments are formed only on lagging strand.
Question 4.
Cytosine is common for both DNA and RNA.
Answer:
It is correct.
Question 5.
RNA does not have guanine as nitrogenous base.
Answer:
Thymine
Question 6.
The complementary base of adenine in RNA molecule is thymine.
Answer:
Uracil
Question 7.
The process of formation of RNA from DNA is translation.
Answer:
Transcription
Question 8.
DNA polymerase-I is mainly responsible for synthesis of new strand during DNA replication.
Answer:
DNA Polymerase
Question 9.
The genetic information from DNA transferred to ribosomes through ribosomal RNA.
Answer:
messenger
Question 10.
The initiation codon AUG normally codes for formylated cystine.
Answer:
methionine
Question 11.
CCC is the initiation codon.
Answer:
AUG
Question 12.
The split genes are needed constantly for cellular activity.
Answer:
housekeeping
Question 13.
The lac operon consists of four regulatory genes only.
Answer:
three
Question 14.
A regulated unit of genetic material for prokaryotic gene expression is called operon.
Answer:
It is correct.
Question 15.
64 codons code for all the 20 essential amino acids.
Answer:
61 codons
Question 16.
Prokaryotic /nRNA is monocistronic.
Answer:
polycistronic
Question 17.
The structural genes are regulated as a unit by a single regulator in operon.
Answer:
promoter
Question 18.
Galactose is an inducer molecule.
Answer:
Lactose
Question 19.
tRNA carries the codes for amino acid sequence.
Answer:
It is correct.
![]()
Fill in the blanks
Question 1.
Frederick Griffith discovered the phenomenon called …………… .
Answer:
transformation
Question 2.
The two strands of polynucleotides forming DNA are ……….. and antiparallel.
Answer:
complementary
Question 3.
A ……………. is located towards 3’ end of the coding strand.
Answer:
OH
Question 4.
The correspondence between triplets in DNA (or RNA) and amino acids in protein is known as ……….. .
Answer:
genetic code
Question 5.
……… is a short sequence of DNA where the repressor binds, preventing RNA polymerase from attaching to the ……….. .
Answer:
Operator, promoter
Question 6.
DNA fingerprinting works on the principle of …………. in DNA sequences.
Answer:
polymorphism
Question 7.
RNA can give rise to DNA through the enzyme ……………. .
Answer:
reverse travscriptase
Question 8.
The movement of a ribosome from 5′-3′ end of mRNA to recognise all codons during protein synthesis is called ………… .
Answer:
elongation
Express in one or two word(s)
Question 1.
The enzyme which joins Okazaki fragments to form a continuous DNA molecule.
Answer:
Ligase
Question 2.
The organism on which Meselson and Stahl (1958) provided strong evidence for semiconservative mode of DNA replication?
Answer:
E. coli
Question 3.
The strand which is transcribed into mRNA (RNA transcript).
Answer:
Template strand
Question 4.
The scientist who formulated central dogma of molecular biology in 1958?
Answer:
Crick
Question 5.
The first X-ray diffraction pattern of DNA was given by which scientist?
Answer:
Wilkins
Question 6.
The scientist who suggested that the genetic code should be made of a combination of three nucleotides.
Answer:
George Gamow
Question 7.
The codon which acts as initiation codon and also codes for amino acid methionine.
Answer:
AUG
Question 8.
All terminator codons begin with nucleotide of which base?
Answer:
U
Question 9.
The scientist who proposed the operon concept?
Answer:
Jacob and Monod
![]()
Short Answer Type Questions
Question 1.
Write a short note on nitrogenous bases.
Answer:
Nitrogenous bases are heterocyclic compounds in which the rings contain both nitrogen and carbon atoms.
There are two types of nitrogenous bases
- Purines (with double rings) adenine and guanine
- Pyrimidines (with single ring) cytosine, uracil and thymine.
Out of the pyrimidines, cytosine is common for both DNA and RNA while thymine is present only in DNA. Uracil is present in RNA in place of thymine.
Question 2.
If a double-stranded DNA has 20% of cytosine, calculate the percentage of adenine in the DNA.
Answer:
Given, cytosine = 20%
∴ Percentage of guanine = 20%
Now according to Chargaff’s rule,
A + T = 100 – (G + C)
⇒ A + T =100 – 40
∴ Percentage of thymine = Percentage of adenine
= \(\frac{60%}{2}\) = 30%
Question 3.
The base sequence in one of the strands of DNA is TAGCATGAT.
(i) Give the base sequence of the complementary strand.
(ii) How are these base pairs held together in a DNA molecule?
(iii) Explain the base complementarity rule. Give the name of the scientist who framed this rule?
Answer:
(i) ATCGTACTA
(ii) Base pairs are held together by weak hydrogen bonds, adenine pairs with thymine by two H-bonds and guanine pairs with cytosine forming three H-bonds.
(iii) According to base complementarity rule formed by Erwin Chargaff for a double-stranded DNA, the ratios between adenine-thymine and guanine-cytosine are constant and equal to one.
Question 4.
A DNA segment has a total of 1000 nucleotides, out of which 240 of them are adenine containing nucleotides. How many pyrimidine bases this DNA segment possesses?
Answer:
According to ChargafFs rule, ratio of purines to pyrimidines is equal, i.e. A + G = C + T
Since, the number of adenine (A) is equal to the number of thymine (T) and A = 240 (given)
Therefore, T = 240
Also, the number of guanine (G) is equal to cytosine (C)
Thus, G+C = 1000 – (A + T)
G + C = 1000-480 = 520
Hence, G = 260, C = 260
The number of pyrimidine bases, i.e.
C + T = 240 + 260 = 500
![]()
Question 5.
It is established that RNA is the first genetic material. Explain giving reasons.
Answer:
RNA is the first genetic material because
- It is capable of both storing genetic information and catalysing chemical reactions.
- Essential life processes such as metabolism, translation, splicing, etc., have evolved around RNA.
- It can directly code for protein synthesis and hence, can easily express the character.
Question 6.
Write short note on RNA.
Answer:
RNA is a genetic material in some viruses.
RNA differs from DNA in having uracil in place of thymine and most RNAs are single-stranded.
It is of three main types, i.e. messenger RNA (mRNA), transfer RNA (tRNA) and ribosomal RNA (rRNA).
mRNA associates with ribosomes for protein synthesis, tRNA is helical and transfers specific amino acids from the cytoplasm to site of protein synthesis while rRNA is part of ribosomes.
Question 7.
Write a short note on tRNA.
Answer:
- tRNA is the smallest form of RNA and functions as in transfering amino acids from cytoplasm to the ribosomes at the time of protein synthesis.
- tRNA has a secondary structure like clover leaf. But its three dimensional structure depicts it as an inverted L-shaped molecule.
- tRNA has five arms or loops, i.e. anticodon loop, amino acid acceptor end, T-loop, D-loop and variable loop.
- tRNAs are specific for specific amino acid.
Question 8.
Explain the two factors responsible for conferring stability to double helix structure of DNA.
Answer:
The factors responsible for stability of double helix structure of DNA are as follows
- Stacking of one base pair over other.
- H-bond between nitrogenous bases.
Question 9.
Which property of DNA double helix led Watson and Crick to hypothesise semiconservative mode of DNA replication? Explain.
Answer:
In the double helical structure of DNA, the two strands of DNA have complementary base pairing and run in opposite direction. This property of DNA double helix led Watson and Crick to hypothesise the semiconservative mode of DNA replication.
Question 10.
Name a few enzymes involved in DNA replication other than DNA polymerase and ligase. Name the key functions for each of them.
Answer:
The enzymes involved in DNA replication other than DNA polymerase and ligase are listed below with their functions
- Helicase – Opens the helix
- Topoisomerases – Removes the tension caused due to unwinding
- DNA ligase – Joins the cut DNA strands
Question 11.
State the dual role of deoxyribonucleoside triphosphates during DNA replication.
Answer:
- The deoxyribonucleoside triphosphates are the building blocks for the DNA strand (polynucleotide chain) is as substrate.
- These also serve as energy source in the form of ATP and GTP from two terminal phosphates.
Question 12.
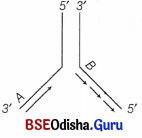
Why do you see two different types of replicating strands in the given DNA replication fork? Explain. Name of these strands.
Answer:
Both the parent strands function as template strands.
On the template strand with 3′ → 5′ polarity, the new strand is synthesised as a continuous strand.
The DNA polymerase can carry out polymerisation of the nucleotides only in 5′ → 3′ direction. This is called continuous synthesis and the strand is called leading strand.
On the other template strand with 5′ → 3′ polarity, the new strand is synthesised from the point of replication fork, also in 5′ → 3′ direction. But, in short fragments, they are later joined by DNA ligases to form a strand called lagging strand.
Question 13.
Write a short note on centrol dogma.
Answer:
Central Dogma
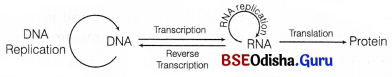
It was proposed by Francis Crick (in 1958). According to the central dogma in molecular biology, the flow of genetic information is unidirectional,’ i.e.
DNA → RNA → Protein.

Central dogma
But later in 1970, HM Temin reported that the flow of information can be in reverse direction also, i.e. from RNA to DNA in some viruses (e.g. HIV) which is called as reverse transcription.
Question 14.
Briefly describe transcription in prokaryotes.
Answer:
Prokaryotes
All three RNAs are needed for synthesis of a protein in a cell. DNA dependent RNA polymerase is the single enzyme that catalyses the transcription of all types of bacterial RNA. But for the expression of different genes, different sigma factors may associate with same core enzymes.
In E.coli, σ70 is used in normal condition σ32 / σH under heat shock, σ54 / σN under nitrogen starvation and σ28 for chemotaxis.
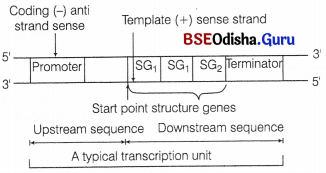
A typical bacterial transcription unit
Question 15.
Describe the initiation process of transcription in bacteria.
Answer:
Initiation
- The holoenzyme binds to the promoter region of transcription unit.
- The sigma polypeptide binds loosely to the promoter sequences so as to form a loose, closed, binary complex.
- It is followed by the formation of a transcription eye or bubble due to the denaturation of adjacent sequence of DNA, lying next to the complex.
- The transcription bubble along with the bounded holoenzyme is called open binary complex.
- In 90% of cases, the start point of transcription is a purine.
- At the elongation site of enzyme, two nucleotides complementary to the first two nucleotides of template strand binds.
- A phosphodiester bond is formed between these two ribonucleotides.
- At this stage, the complex is called ternary complex that consists of partly denatured DNA bounded with holoenzyme having a di-ribonucleotide.
- The same process continues till a RNA chain of about nine nucleotides is synthesised. The holoenzyme does not move throughout this process.
- After the completion of initiation process, sigma factor dissociates from RNA polymerase. This facilitates the promoter clearance so that a new holoenzyme can bind to promoter for second round of transcription.
Question 16.
Explain (in one or two lines) the function of the following
(i) Introns
(ii) Exons
Answer:
The primary transcript contains both the exons and introns and these are non-functional.
Such genes are called split genes/interrupted genes. Therefore, they undergo a process called splicing to remove the introns and to join the exons in a proper order to allow translation to take place.
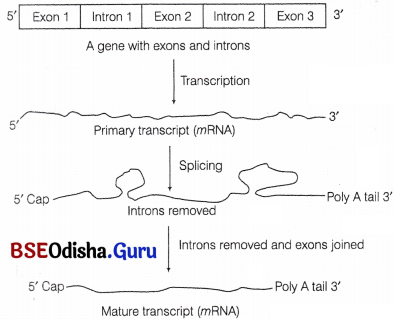
Expression of an interrupted/split gene and RNA splicing
The presence of introns is reminiscent of antiquity and the process of splicing represents the dominance of RNA world. The bnRNA undergoes two additional processes, i. e. post-transcriptional modifications.
Question 17.
State the difference between the structural genes in a transcription unit of prokaryotes and eukaryotes.
Answer:
Prokaryotic structural genes are found in continuity without any non-coding region, while eukaryotic structural genes are divided into exons (coding DNA) and introns (non-coding DNA). Exons appear in mature RNA. Introns are spliced out during splicing.
Question 18.
Depending upon the chemical nature of the template (DNA or RNA) and the nature of nucleic acids synthesised from it (DNA or RNA), list the types of nucleic acid polymerases.
Answer:
- DNA-dependent DNA polymerase uses DNA template to catalyse the polymerisation of deoxynucleotides.
- DNA-dependent RNA polymerase catalyses transcription of all types of RNAs in bacteria.
- In eukaryotes, there are three types of DNA-dependent RNA polymerase
- RNA polymerase-I transcribes rRNAs.
- RNA polymerase-II transcribes precursor of mRNA.
- RNA polymerase-III transcribes tRNA, srRNA and snRNAs.
Question 19.
Why hnRNAis required to undergo splicing?
Answer:
hnKNA is required to undergo splicing because of the presence of introns (the non-coding sequences) in it. These need to be removed and the exons (the coding sequences) have to be joined in a specific sequence for translation to take place.
Question 20.
Write a note on genetic code.
Answer:
Francis Crick conducted an experiment in Viral DNA in 1961 and concluded that genetic code is triplet and with any punctuation it is read continuously. Once the triplet nature of codon was established, different scientists then tried to establish codons for 20 different amino acids found in proteins.
- Marshall Nirenberg In 1961, he used a synthetic twRNA of uracil only. He found that the translated polypeptide was composed of amino acid-phenylalanine only Thus, he concluded the UUU codon codes for phenylalanine.
- Nirenberg and Philip Leder In- 1964, they found 47 out of 64 possible codons by employing the technique of triplet binding.
- Har Gobind Khorana He worked out remaining 17 codons by employing artificial bzRNA.
Note In 1968, Nirenberg and Khorana shared Nobel Prize with RW Holley (gave details of fRNA structure).
On the basis of above discoveries, the spellings of the genetic code were put together in a checkerboard, as given below
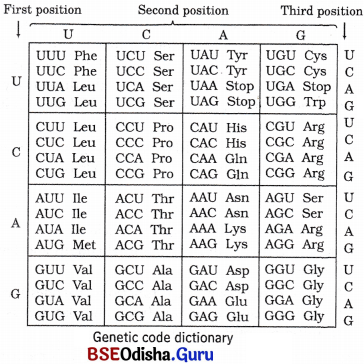
Question 21.
Write short note on peptide bonds.
Answer:
Amino acids are joined together in proteins by peptide bonds. A peptide bond is a covalent bond formed between two molecules when the carboxyl group of one molecule reacts with the amine group of the other molecule. This releases a water molecule. CONH is called a peptide link.
Question 22.
Unambiguous, universal and degenerate are some of the terms used for the genetic code. Explain the salient features of each of them.
Answer:
Unambiguous code means that one codon codes for only one amino acid, i.e. AUG codes only for methionine. Genetic code is universal, as particular codon codes for the same amino acid in all organisms. It is degenerate because some amino acids are coded by more than one codon, e.g. UUU and UUC, both code for phenylalanine.
Question 23.
Write short note on aminoacylation in translation.
Ans.
Amino acids in the cytoplasm are inactive. They cannot take part directly in protein synthesis or translation. The formation of peptide bond requires energy. In the presence of ATP, amino acids become activated by binding with aminoacyl fRNA synthase enzyme, i.e. aminoacylation.
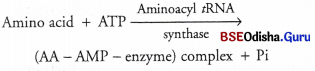
Question 24.
Write short note on operon.
Or
Write a note on operon concept.
Answer:
An operon is a unit of prokaryotic gene expression which includes sequentially regulated structural genes and control elements recognised by the regulatory gene product. F Jacob and J Monod gave the operon concept and were the first ones to describe a transcriptionally regulated system.
Question 25.
What is DNA fingerprinting? Mention its applications.
Or Write a note on DNA fingerprinting.
Answer:
DNA Fingerprinting:
The technique of DNA fingerprinting or DNA typing or DNA profiling was developed and established by British geneticist Dr. Alec Jeffreys based on the fact that like every individual organism is unique in its fingerprints. The DNA pattern also differs in every individual.
Fingerprints can be altered by surgery but there is no known procedure available to alter the DNA design of an individual. For obtaining the DNA fingerprints of an individual, highly polymorphic genes that occur in multiple forms in different individuals are selected.
Applications of DNA Fingerprinting:
This technique can he applied in various fields such as
- Used as a tool in forensic investigations.
- To settle paternity disputes.
- To study evolution by determining the genetic diversities among population.
Question 26.
(i) Explain DNA polymorphism as the basis of genetic mapping of human genome.
(ii) State the role of VNTR in DNA fingerprinting.
Answer:
(i) Polymorphism is inherited from parents to children. So, it is useful for the identification (forensic application) and paternity testing. It arises due to mutations and also plays an important role in evolution and speciation. These mutations in the non-coding sequences have piled up with time and form the basis of DNA polymorphism. It is the basis of genetic mapping of human genome as well as DNA fingerprinting.
(ii) Variable Number of Tandem Repeats (VNTRs) belong to a class of satellite DNA called as minisatellite. VNTR are used as probes in DNA fingerprinting.
Long Answer Type Questions
Question 1.
Describe the structure of DNA with a neat and labelled diagram.
Or Describe the structure of DNA molecule as per the model proposed by Watson and Crick.
Answer:
Primary Structure of DNA
Two nucleotides when linked through a 3′ → 5′ phosphodiester linkage, form a dinucleotide. The phosphodiester linkage is formed when each phosphate group esterifies to the 3′ hydroxy! group of a pentose and to the 5′ hydroxyl group of the next pentose.
In a similar fashion, more nucleotides may join to form a polynucleotide chain (fig. structure of DNA). The polymer chain thus, formed has
- One end with a free phosphate moiety at 5′ end of deoxyribose sugar. This is marked as 5′ end of polynucleotide chain.
- The other end with a free hydroxyl 3′ – OH group marked as 3′ end of the polynucleotide chain.
Thus, the sugar and phosphates form the backbone in a polymer chain and the nitrogenous bases linked to sugar moiety project from this backbone. In RNA, there is an additional – OH group at 2′ position in the ribose of every nucleotide residue.
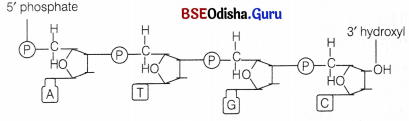
A Ploynucleotide chain
Secondary Structure of DNA:
Watson and Crick proposed the secondary structure in the form of the famous double helix model in 1953 on the basis of following observations
1. Erwin Chargafif (in 1950) formulated important generalisation on the base and other contents of DNA, called as ChargafFs rule. It states that for a double-stranded DNA, the ratios between adenine (A) and thymine (T) and guanine (G) and cytosine (C) are constant and equal to one.
i.e. \(\frac{A+T}{G+C}\) = 1
2. X-ray diffraction studies by Wilkins in 1952, suggested a helicoidal configuration of DNA.
One of the important features of this model was the complementary base pairing. It means if the sequence of bases in one strand is known, the sequence in other strand can be easily predicted. Also, if each strand from a DNA acts as a template for synthesis of a new strand, the daughter DNA thus produced would be identical to the parental DNA molecule.
Watson and Crick Model of DNA:
Watson and Crick worked out the first correct double helix model of DNA, which explained most of its properties.
The salient features of double helix structure of DNA are as follows
1. DNA is made up of two polynucleotide chains. The backbone is constituted by sugar phosphate, while the nitrogenous bases project inwards.
2. The two chains have anti-parallel polarity, i.e. when one chain has 3′ → 5′ polarity, the other has 5′ → 3′ polarity. Hence, orientation of deoxyribose sugar is opposite in both the strands.
3. The two strands are complementary to each other, i.e. purine base of one strand has pyrimidine counterpart on other strand. The complementary bases in two strands are paired through hydrogen bonds (H-bonds) to form base pairs.
(a) Adenine is bonded with thymine of the opposite strand with the help of two hydrogen bonds.
(b) Guanine is bonded with cytosine of the opposite strand with the help of three hydrogen bonds. So, a purine bonds with a pyrimidine always. Thus, maintaining a uniform distance between the two strands of the helix.
4. The two polypeptide chains are coiled in a right-handed fashion. Pitch of the helix, i.e. length of DNA in one complete turn = 3.4 nm or
3.4 × 10-9 or 34 Å.
Number of base pairs in each turn = 10. Distance between a base pair in a helix = 0.34 nm. The diameter of DNA molecule is 20 Å (2nm).
5. Percentage calculation of bases is done by A + T = 100 – (G + C).
6. The plane of one base pair stacks over the other in double helix. This provides the stability to the helical structure, in addition to H-bond.
The length of DNA in E. coli is 1.36 mm, while in humans it is 2.2 m.
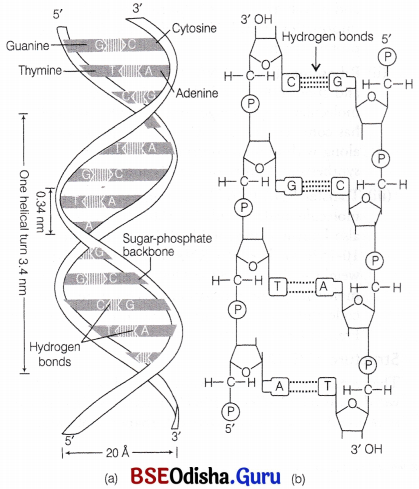
Structure of DNA : (a) Watson and Crick model of double helix, (b) Double-stranded polynucleotide chain sequence showing hydrogen bonds
Question 2.
Give an account of Griffith’s experiment on transformation.
Answer:
Frederick Griffith in 1928, carried out a series of experiments with Diplococcus pneumoniae (a bacterium that causes pneumonia). He observed that when these bacteria were grown on a culture plate, some of them produced smooth, shiny colonies (S-type), whereas the others produced rough colonies (R-type).
This difference in appearance of colonies (smooth/rough) is due to the presence of mucous (polysaccharide) coat on S-strains (virulent/pathogenic) but not on R-strains (avirulent/non-pathogenic).
Experiment:
- He first infected two separate groups of mice. The mice that were infected with the S-strain (S-III) died from pneumonia as S-strains are the virulent strains causing pneumonia.
- The mice that were infected with the R-strain (R-II) did not develop pneumonia and they lived.
- In the next set of experiments, Griffith killed the bacteria by heating them. The mice that were injected with heat-killed S-strain bacteria did not die and lived.
- Whereas, on injecting a mixture of heat-killed S-strain and live R-strain bacteria, the mice died. Moreover, living S-bacteria were recovered from the dead mice.
These steps are summarised below

From all these observations Griffith concluded that the live R-strain bacteria, had been transformed by the heat-killed S-strain bacteria, i.e. some ‘transforming principle’ had transferred from the heat-killed S-strain, which helped the R-strain bacteria to synthesise a smooth polysaccharide coat and thus, become virulent.
This must be due to the transfer of the genetic material. However, he was not able to define the biochemical nature of genetic material from his experiments.
Biochemical Characterisation of Transforming Principle:
Oswald Avery, Colin MacLeod and Maclyn McCarty (1933-44) worked in Rockfellar Institute, New Xork, USA to determine the biochemical nature of ‘transforming principle’ in Griffith’s experiment in an in vitro system. Prior to this experiment, the genetic material was thought to be protein.
During this experiment, purified biochemicals (i.e. proteins, DNA, RNA, etc.) from the heat-killed S-III cells were taken, to observe which biochemicals could . transform live R-cells into S-cells.
They discovered that DNA alone from heat-killed S-type bacteria caused the transformation of non-virulent R-type bacteria into S-type virulent bacteria.
They also discovered that protein digesting enzymes (proteases) and RNA digesting enzymes (ribonuclease) did not inhibit this transformation. This proved that the ‘transforming substance’ was neither protein nor RNA.
DNA-digesting enzyme (deoxyribonuclease) caused inhibition of transformation, which suggests that the DNA caused the transformation. This provided the first evidence for DNA as transforming principle or the genetic material.
The steps of this experiment are summarised below
- R-II + DNA extract of S-III + no enzyme = R-II colonies + S-III colonies
- R-II + DNA extract of S-III + Ribonuclease = R-II colonies + S-III colonies
- R-II + DNA extract of S-III + Protease = R-II colonies + S-III colonies
- R-II + DNA extract of S-III + Deoxyribonuclease = Only R-II colonies
Question 3.
State the aim and describe Meselson and Stahl’s experiment.
Answer:
Meselson and Stahl in 1958, aimed to prove that DNA replicates in a semiconservative fashion. The semiconservative DNA replication suggests that, after the completion of replication, each DNA molecule will have one parental and one newly synthesised strand.
Meselson and Stahl’s experiment
Matthew, Meselson and Franklin Stahl in 1958 conducted an experiment in California Institute of technology with Escherichia coli to prove that DNA replicates semiconservatively as follows
1. They grew many generations of E.coli in a medium containing 15NH4Cl (15N is the heavy isotope of nitrogen) as the only source of nitrogen.
The result was that 15N got incorporated into the newly synthesised DNA after several generations. By centrifugation in a cesium chloride (CsCl) density gradient, this heavy DNA molecule could be distinguished from the normal DNA.
2. The bacterial culture was then washed to make the medium free and the cells were then transferred into a medium containing normal 14NH4Cl.
3. The samples were separated independently on CsCl gradient to measure the densities of DNA.
4. At definite time intervals, as the cells multiplied, samples were taken and the DNA which remained as double-stranded helices was extracted.
5. The DNA obtained from the culture, one generation after the transfer from 15N to 14N medium (i.e. after 20 minutes because E.coli divides in 20 minutes) had a hybrid or intermediate density.
This hybrid density was the result of replication in which DNA double helix had separated and the old strand (N15) had synthesised a new strand (N14).
6. The DNA obtained from the culture after another generation (II generation) was composed of equal amounts of hybrid DNA containing (N15 molecule) and ‘light’ DNA (containing 14N molecule).
7. With the preeceding growth generations in normal medium, more and more light DNA was present. Hence, the semiconservative mode of DNA replication was confirmed.
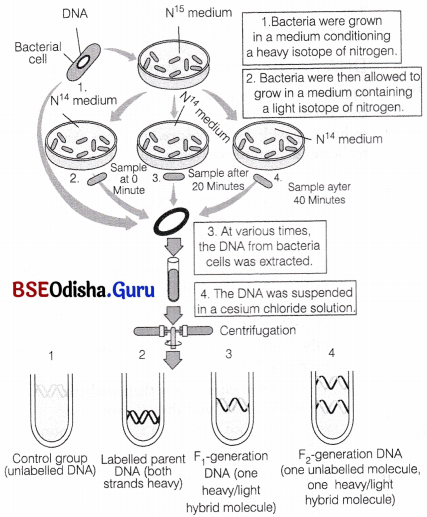
Meselson-Stahl experiment to demonstrate semiconservative replication
Similar experiments on a eukaryote, ‘ Vicia faba’ (faba beans) were conducted by Taylor and colleagues in 1958, involving use of radioactive thymidine, i.e. bromodeoxy-uridine. The replicated chromosomes were then, stained with fluorescent dye and Giemsa. The old and new strand were stained differently which confirm that the DNA in chromosomes also replicates semiconservatively.
Question 4.
Describe the process of DNA replication.
Answer:
DNA Replication:
In addition to the double helical structure of DNA, Watson and Crick also proposed a scheme for DNA replication. According to this model, the two strands of double helix separate and act as a template for the synthesis of new complementary strands in which the base sequence of one strand determines the sequence on the other strand.
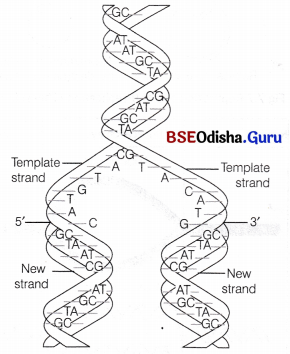
Watson and Crick model for semiconservative DIMA replication
This is called base complementarity and it ensures the accurate replication of DNA. After the completion of replication, each DNA molecule have one parental and one newly synthesised strand.
This scheme for DNA replication was termed as semiconservative DNA replication.
DNA Replication is Semiconservative:
Meselson and Stahl in 1958, aimed to prove that DNA replicates in a semiconservative fashion. The semiconservative DNA replication suggests that, after the completion of replication, each DNA molecule will have one parental and one newly synthesised strand.
Meselson and Stahl’s experiment
Matthew, Meselson and Franklin Stahl in 1958 conducted an experiment in California Institute of technology with Escherichia coli to prove that DNA replicates semiconservatively as follows
1. They grew many generations of E.coli in a medium containing 15NH4Cl (15N is the heavy isotope of nitrogen) as the only source of nitrogen.
The result was that 15N got incorporated into the newly synthesised DNA after several generations. By centrifugation in a cesium chloride (CsCl) density gradient, this heavy DNA molecule could be distinguished from the normal DNA.
2. The bacterial culture was then washed to make the medium free and the cells were then transferred into a medium containing normal 14NH4Cl.
3. The samples were separated independently on CsCl gradient to measure the densities of DNA.
4. At definite time intervals, as the cells multiplied, samples were taken and the DNA which remained as double-stranded helices was extracted.
5. The DNA obtained from the culture, one generation after the transfer from 15N to 14N medium (i.e. after 20 minutes because E.coli divides in 20 minutes) had a hybrid or intermediate density.
This hybrid density was the result of replication in which DNA double helix had separated and the old strand (N15) had synthesised a new strand (N14).
6. The DNA obtained from the culture after another generation (II generation) was composed of equal amounts of hybrid DNA containing (N15 molecule) and ‘light’ DNA (containing 14N molecule).
7. With the preeceding growth generations in normal medium, more and more light DNA was present. Hence, the semiconservative mode of DNA replication was confirmed.

Meselson-Stahl experiment to demonstrate semiconservative replication
Similar experiments on a eukaryote, ‘ Vicia faba’ (faba beans) were conducted by Taylor and colleagues in 1958, involving use of radioactive thymidine, i.e. bromodeoxy-uridine. The replicated chromosomes were then, stained with fluorescent dye and Giemsa. The old and new strand were stained differently which confirm that the DNA in chromosomes also replicates semiconservatively.
Pre-Requisites of DNA Replication:
DNA replication is a complex process that requires many enzymes and protein factors. This process is very fast and accurate. It is seen in both prokaryotes and eukaryotes and involves some basic steps that are listed below
(i) Two parental strands unwind and get separate.
(ii) One of the parental strand acts as a template for the synthesis of new strand. Hence, a new strand is formed by the winding of one old and one new strand.
Major components involved in the process of DNA replication are as follows
I. On (Origin of Replication)
- It is the specific site on DNA where replication starts and proceeds in one or both directions. This site is known as origin of replication (Ori).
- Ori specifying DNA segments can be isolated from E. coli, Coli phages, plasmids, yeasts and eukaryotic viruses.
- Ori in E. coli is called Ori C. It is a DNA sequence of about 245 base pairs that is rich in A-T bases. Hence, the two strands easily get separated at the origin.
- Ori in Yeast is called Autonomous Replication . Sequence (ARS) which is 150 base pair long. It acts as the binding site for Origin Recognition Complex (ORC).
- During replication, Ori is recognised by replication initiator complex and the process of replication starts. It proceeds along the replication forks.
- Each Ori has two termini. A replicon is one Ori (or origin) with its two unique termini. In prokaryotes (E.coli), the entire circular DNA acts as a single replicon. In eukaryotes, DNA is larger and hence, they have several Ori per DNA.
- In bidirectional replication, the two strands separate at the origin or Ori. It results in the formation of a replication eye and both ends move along the replication, e.g. θ-replication (Theta) in prokaryotes.
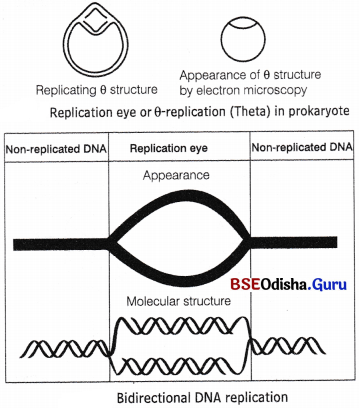
- In unidirectional replication, one of the two ends of the replication eye moves along the replication fork while the other end remains stationary, e.g. replication of mitochondrial DNA (mt DNA) in vertebrates.
II. DNA Polymerase
- It is the main enzyme which uses a DNA template to catalyse the polymerisation of deoxynucleotides. The average rate of polymerisation is 2000 bp (base pairs) per second approximately.
- DNA polymerase adds deoxyribonucleotides to the 3′-OH end of polynucleotide by the removal of pyrophosphate from nucleoside triphosphate.
- Polynucleotide (n) + d NTP → Polynucleotide (n + 1) + PPi
Types of DNA Polymerases
- In prokaryotes, there are three types of DNA polymerases, i.e. DNA polymerase-I, II and III, whereas in eukaryotes, five different DNA polymerases have been indentified, i.e. DNA polymerases α, ß, γ, δ and ε.
- DNA polymerase was first isolated from Exoli by Arthur Kornberg in Washington University in 1956. It was first called Kornberg enzyme but later its name was changed to DNA polymerase-I due to the discoveries of other polymerases.
- DNA polymerase-I and II Involved in DNA repair and proofreading in prokaryotes.
- DNA polymerase-III It has exonuclease activity. It can remove nucleotides from 3′ end of DNA strand, i.e., 3’→5′ exonuclease. It also helps in proofreading so that wrong nucleotide added at 3′ end can be removed.
- Exonuclease activity of different DNA polymerases is listed below
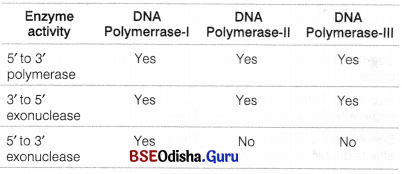
Working of DNA Polymerase
DNA polymerase (Pol) requires a template to synthesise a new strand in 5′-3′ direction. For this, they add a primer to the template strand. The new nucleotides are then added to the 3′-OH end of primer so that the synthesis of DNA proceeds in 5′-3′ direction.
Note A prime is small DNA or RNA strand that is to template strand through hydrogen bonds.
III. Other Enzymes
Besides DNA polymerases, other enzymes involved in the process of DNA replication are as follows
- Helicase It unwinds the DNA strand, i.e. separates the two strands from one point, for the formation of a
replication fork. - Topoisomerase (DNA gyrase) The unwinding of DNA creates a tension in the DNA strands, which get released by the enzyme topoisomerase.
- DNA Ligase It facilitates the joining of DNA strands together by catalysing the formation of phosphodiester bond. It plays a role in repairing single-strand breaks in duplex DNA.
Mechanism of DNA Replication:
All the enzymes and protein factors involved in DNA replication constitutes a replicase system or replisome. . The process of replication proceeds in the following steps
- An initiator protein recognises the Origin of replication (Ori) and binds to it.
- DNA helicase enzyme breaks the hydrogen bonds between nitrogenous bases and unwinds the ds DNA.
- To prevent rewinding and attack by single stranded nuclease, Single-Stranded Binding proteins (SSB proteins) bind to the separated strands. SSB also help to keep these ssDNA in extended position.
- The combined action of helicase and SSB proteins results in the formation of V-shaped replication fork at the origin.
- As the replication fork moves, DNA unwinds and a positive super coil is formed in the unreplicated portion of DNA, i.e. in front of the fork.
- The super coil is like a knot that hinders the fork movement. It is removed by the enzyme topoisomerase (gyrase) in E.coli and topoisomerase-I in eukaryotes.
Question 5.
How do mRNA, tRNA and ribosomes help in the process of translation?
Answer:
(i) Binding of mRNA to Ribosome:
Ribosomes occur in the cytoplasm as separate smaller and larger units.
In prokaryotes, IF-3 binds to 30 S subunit to prevent association of subunits. The incoming tRNA containing specific amino acid binds to the A-site. The prokaryotic mRNA has a leader sequence called shine-Delgarno sequence that is present just prior to initiation codon-AUG.
It is homologous to 3′ end of 16S rRNA (ASD region) in 30S subunit. This complementarity ensures correct binding of 30S subunit on mRNA. The peptidyl tRNA containing elongating polypeptide then binds to P-site. The bacterial ribosome contains another site, the E-site or Exit-site to which the discharged tRNA or tRNA whose peptidyl has already been transferred binds before its release from ribosome.
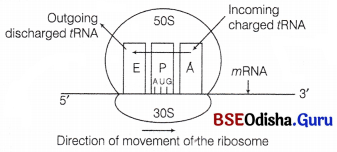
A bacterial ribosome with three sites; E, exit site, P, peptidyl site and A, aminoacyl site
(ii) Aminoacylation
Aminoacylation of tRNA The activation of amino acids takes place through their carboxyl groups. The amino acids are activated in the presence of ATP and linked to their cognate tRNA. In the presence of ATP, amino acids become activated by binding with aminoacyl tRNA synthetase enzyme.
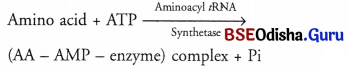
The amino acid AMP-enzyme or AA-AMP enzyme complex is called an activated amino acid.
In the second step, this complex associated with the 3′-OH end of tRNA. AMP gets hydrolysed to form an ester bond between amino acid and tRNA and the enzyme is released.
AA – AMP – enzymes + tRNA → AA – tRNA + AMP + enzyme.
(iii) Transfer of Activated Amino Acids to tRNA
The amino acids are attached to the tRNA by high energy bonds. These bonds are formed between the carboxyl group of amino acid and 3′ hydroxy terminal of ribose of terminal adenosine of CCA and of tRNA.
The complete reaction is carried out by enzyme synthetase which has two active sites, i.e. one for tRNA and another for specific amino acid molecule.
Question 6.
Describe the process of translation in prokaryotes.
Or Describe the initiation step of translation in prokaryotes.
Answer:
Translation requires a machinery which consists of ribosome, wzRNA, rRNAs, aminoacyl rRNA synthetase (enzyme that helps in combining amino acid to particular rRNA) and amino acids.
Initiator tRNA
It is a specific rRNA for the process of initiation and there are no rRNAs for stop codons.
Ribosome
It occurs in cytoplasm and responsible for protein synthesis. It consists of structural RNAs and around 80 different proteins. Ribosome exists as two subunits in its inactive stage
(i) Small subunit When the small subunit encounters an mRNA, translation of mRNA to protein begins.
(ii) Large subunit It consists of two sites where amino acids can bind to and be close to each other for the formation of a peptide bond. Ribosome also acts as a catalyst (23S rRNA in bacteria is the enzyme ribozyme) for peptide bond formation.
Translational Unit:
It is the sequence of RNA flanked by the start codon (AUG) and the stop codon in mRNA. It codes for a polypeptide that has to be produced.
Untranslated Regions (UTR):
These are some additional sequences in an wRNA that are not translated. They are present at both the ends, i.e. at 5′ end (before start codon) and at 3′ end (after stop codon). They improve the efficiency of translation process.
Mechanism of Translation:
The main steps in translation include
(i) Binding of mRNA to ribosome
(ii) Activation of amino acids (aminoacylation of tRNA).
(iii) Transfer of activated amino acids to tRNA.
(iv) Initiation of polypeptide chain synthesis.
(v) Elongation of polypeptide chain.
(vi) Termination of polypeptide chain formation.
(i) Binding of mRNA to Ribosome:
Ribosomes occur in the cytoplasm as separate smaller and larger units.
In prokaryotes, IF-3 binds to 30 S subunit to prevent association of subunits. The incoming tRNA containing specific amino acid binds to the A-site. The prokaryotic mRNA has a leader sequence called shine-Delgarno sequence that is present just prior to initiation codon-AUG.
It is homologous to 3′ end of 16S rRNA (ASD region) in 30S subunit. This complementarity ensures correct binding of 30S subunit on mRNA. The peptidyl tRNA containing elongating polypeptide then binds to P-site. The bacterial ribosome contains another site, the E-site or Exit-site to which the discharged tRNA or tRNA whose peptidyl has already been transferred binds before its release from ribosome.

A bacterial ribosome with three sites; E, exit site, P, peptidyl site and A, aminoacyl site
(ii) Aminoacylation
Aminoacylation of tRNA The activation of amino acids takes place through their carboxyl groups. The amino acids are activated in the presence of ATP and linked to their cognate tRNA. In the presence of ATP, amino acids become activated by binding with aminoacyl tRNA synthetase enzyme.
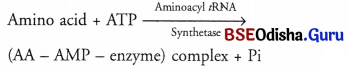
The amino acid AMP-enzyme or AA-AMP enzyme complex is called an activated amino acid.
In the second step, this complex associated with the 3′-OH end of tRNA. AMP gets hydrolysed to form an ester bond between amino acid and tRNA and the enzyme is released.
AA – AMP – enzymes + tRNA → AA – tRNA + AMP + enzyme.
(iii) Transfer of Activated Amino Acids to tRNA
The amino acids are attached to the tRNA by high energy bonds. These bonds are formed between the carboxyl group of amino acid and 3′ hydroxy terminal of ribose of terminal adenosine of CCA and of tRNA.
The complete reaction is carried out by enzyme synthetase which has two active sites, i.e. one for tRNA and another for specific amino acid molecule.
(iv) Initiation of Polypeptide Chain Synthesis:
The protein synthesis begins from the amino terminal end of the polypeptide, proceeds by the addition of amino acids through peptide bond formation and ends at the carboxyl terminal end. In prokaryotes, the initiation amino acid is formylated methionine while in eukaryotes it is methionine.
Initiation in Prokaryotes
In prokaryotes, two types of tRNA are present for methionine
(a) tRNAfmet for initiation carrying formyl methionine and
(b) tRNAmet for carrying normal methionine to growing polypeptide.
The initiation of polypeptide synthesis requires the following components
mRNA, 30S subunit of ribosome, formylmethionyl-tRNA (fmet-tRNAfmet), initiation factors IF-1, IF-2 and IF-3, GTP, 50S ribosomal subunit and Mg+2.
The sequence of events occurring during initiation process are
1. The smaller 30S subunit of ribosome binds to the transcription factor IF-3. It prevents the premature association of two ribosomal subunits.
2. Interaction of SD region of mRNA and ASD region of ribosome helps the mRNA to bind to 30S subunit. It also helps AUG to correctly positioned at the P-site of the ribosome.
3. The fMet-tRNAfmet (the specific tRNA aminoacylated to formyl methionine) binds to the AUG codon at the P-site. The tRNAfmct is the only tRNA that binds to its codon present on the P-site. All other fRNA along with their respective amino acids bind to their codon present at the A-site. Therefore, AUG codon present as initiation codon codes for formylmethionine. When it is present at other position it codes for normal methionine.
4. The initiation factor IF-1, binds to the A-site. It prevents the binding of any other aminoacyl tRNA to the codon at the A-site during initiation.
5. The GTP bound IF-2 (GTP-IF-2) and the initiating f Met-tRNAfmet attaches to the complex of 30S subunit-IF3-IF1-mRNA.
6. 50S subunit then attaches the complex formed in the previous step. The GTP bound to IF-2 is hydrolysed to GDP and Pi. After this step, all the three initiation factors leave ribosome. This complex of 70S ribosome, mRNA and f Met-rRNA fmet bound to initiation codon at P site is known as initiation complex.
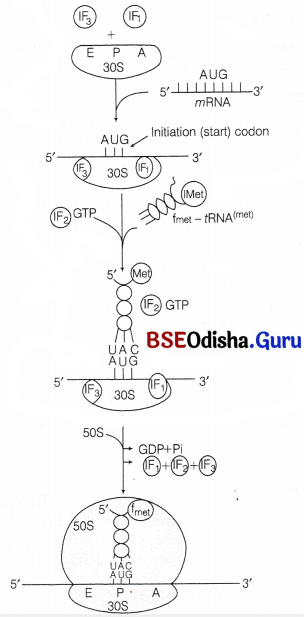
Stepwise formation of initiation complex in prokaryote
(v) Elongation of Polypeptide Chain
In this step, another charged aminoacyl tRNA complex binds to the A-site of the ribosome, following the hydrolysis of GTP to GDP and Pi. A peptide bond forms between carboxyl group (-COOH) of amino acid at P-site and amino group (-NH3) of amino acid at A-site by the enzyme peptidyl transferase.
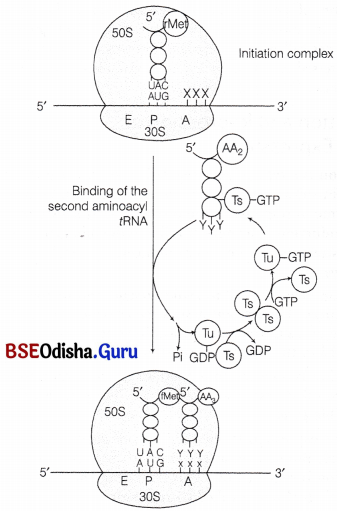
Binding of the second aminoacyl tRNA to the A site of ribosome
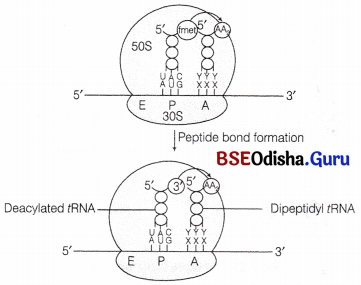
Formation of a peptide bond
(vi) Translocation of Polypeptide:
The peptidyl tRNA bounded to A-site comes to the P-site of ribosome.
The empty tRNA comes to E-site and a new codon occupies the A-site for next aminoacyl tRNA.
- This is achieved by the movement or translocation of ribosome by a codon in 5′ to 3′ direction of mRNA in the presence of EF-G (translocase) and GTP.
- tRNA interact with E-site on 50S subunit through it CCA sequence at 3′ end.
The tRNA molecule is then, transferred from A site to P-site and from P-site to E-site by the movement of two subunits of ribosomes.
Finally, the deacylated tRNA is released to cytosol from E-site.
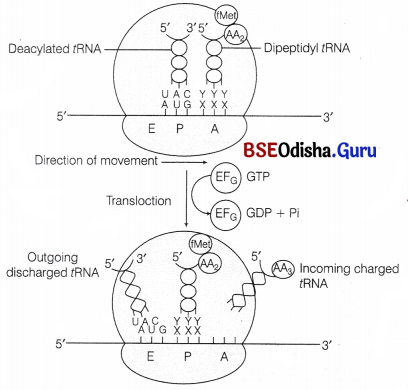
(vii) Termination:
It is accomplished by the presence of any of the three termination codon on mRNA. Which are recognised by the release (termination) factor, RF1, RF2 and RF3.
RF1 and RF2 They resemble the structure of tRNA. They compete with tRNA to bind the termination codon at A-site of ribosome. This is called molecular mimicry. RF1 recognise UAG and RF2 recognise UGA. UAA is recognised by both of them. RF3 helps to split the peptidyl tRNA body by using GTP.
Note In eukaryotes, only one release factor is known. It iseRF1.
